All published articles of this journal are available on ScienceDirect.
The Presence of Microbial Air Contaminants in the Operating Theatre at a Teaching Hospital in East Coast Malaysia
Abstract
Aim:
The aim of this study is to determine the presence of microbial air contaminants in the operating theatre at a teaching hospital.
Objective:
Airborne microbial level in operation theatre is one of the significant risks in hospital as it can increase the surgical site infection and nosocomial infections.
Background:
Duo SAS Super 360 Air Sampler was used to collect the airborne samples in triplicate each for nutrient and MacConkey agar at eight operation rooms and two corridors for morning and evening sessions. Sampling was conducted for three months and repeated every two months. Microbiological culture, gram staining and biochemical tests such as catalase test, oxidase test, coagulase test, Triple Sugar Iron Agar test, urease test, citrate test, Sulfide, Indole, Motility test, Methyl Red Voges-Proskauer test, disc diffusion test, and Albert’s stain were performed on the pure isolated culture.
Methods:
Sampling was conducted for three months and repeated every two months. Microbiological culture, gram staining and biochemical tests such as catalase test, oxidase test, coagulase test, Triple Sugar Iron Agar test, urease test, citrate test, Sulfide, Indole, Motility test, Methyl Red Voges- Proskauer test, disc diffusion test, and Albert’s stain were performed on the pure isolated culture. Bacteria that were present in the operation rooms were Bacillus spp., Micrococcus spp. and Staphylococcus spp. while Pseudomonas aeruginosa and Acinetobacter baumannii were absent. The bacteria identified in the operation theater may also cause surgical site infections and nosocomial infections to the patients, although the microbial contamination in the air of the operation theatre is low.
Results:
The bacteria identified in the operation theater may also cause surgical site infections and nosocomial infections to the patients, although the microbial contamination in the air of the operation theatre is low.
Conclusion:
Strengthening surveillance on the hygienic condition of the operation theatre and routine sampling is strongly recommended to control all possible sources and types of infection.
1. INTRODUCTION
Airborne transmission is one of the routes responsible for the spreading of disease for nosocomial infections [1]. Airborne transmission occurs when infectious agents are carried in the suspended air. Nosocomial infections are also known as hospital-acquired infections. It is an infection that was acquired in the hospital environment, which is not present during the time the patient was admitted [2]. The study of microbial in the indoor air of the hospital is important because it will give more understanding about the bioburden of the airborne microbes in the hospital environment. The hospital is where the patients are treated; hence may influence the health of patients to either recovering or acquiring infections that can lead to death.
Hospital is an important indoor environment responsible for the spread of air pathogens. It serves as a reservoir of pathogens. Individuals can be infected with a number of microorganisms, and then these pathogens are potentially transmitted to other individuals, including patients, hospital workers, and even visitors. Besides, the hospital is referred to as red as a special public place. Moreover, the number of people in the hospital is much higher compared to other public places [2]. In addition to that, one of the areas in the hospital that has a high risk of nosocomial infections among patients in the Operation Theatre (OT) room [3]. This is because the OT room is a place where all the surgeries are conducted. The patients are potentially exposed to the OT environments that may harbor many microbes such as Pseudomonas aeruginosa, Bacillus spp., Acinetobacter baumannii, Micrococcus spp. and methicillin-resistant Staphylococcus aureus (MRSA), which are the major pathogens associated with nosocomial infections [4]. Nosocomial infections in surgical patients are 38%, while Hospital-Acquired Infections (HAI) is ranged between 13% to 40%, respectively, in Italy, Europe, and the United States of America [5].
Surgical Site Infection (SSI) is the second most common infectious complication after urinary tract infection following cesarean section delivery [6]. They reported that in total, 18.8% of their study participants who were among patients undergoing cesarean section developed SSI. Approximately 20-30% of surgical infections are caused by Staphylococcus aureus [7]. Another previous study conducted at Hospital USM (HUSM) in 2015 reported that among 220 patients, the sample collected showed that almost 11.8% incidence was found to be surgical site infections [8].
The OT should be a clean and sterile place from microbes and dust. However, during the study period, the OT was in the process of renovation. Thus, it is hypothesized that renovation may emit foreign particles within the OT room and cause the air in the OT to be contaminated. The foreign particle may lead to nosocomial infections among the patients that already have a weak immune system due to the disease itself, causing them to be highly at risk of SSI. The renovation was conducted for six months; however, the OT was still under operation for surgeries. It is postulated that the microbial level in the indoor air will also be affected due to the renovations. As the effect of the hospital’s indoor air quality to humans is significant, hence it is important to conduct this preliminary study in determining the microbial contamination level at the OT. Moreover, there are limited studies that focused on indoor airborne microbial conducted in hospitals in Malaysia.
2. MATERIALS AND METHODS
This is a cross-sectional study design in isolating bacteria in the OT of a teaching hospital. This study involved sample collection at eight Operation Rooms (ORs) and two corridors from September 2018 to January 2019, in which there was a renovation activity happening in the OT. Sampling was conducted thrice, which was repeated every two months.
2.1. Airborne Microbial Sampling
The airborne microbial level was sampled using a Duo SAS Super 360 (SAS VWR, Italy) with two sampling heads. This study used two types of agar medium, which were Nutrient Agar (NA) to collect airborne microbial such as Bacillus spp., Micrococcus spp. and Staphylococcus spp. while MacConkey Agar (MAC) to collect Pseudomonas aeruginosa and Acinetobacter baumannii. The SAS air sampler was then set to sample with required airflow 1000 ml for about 8 minutes. The SAS air sampler was placed at the height of one meter from the floor in the ORs and corridors.
2.2. Isolation and Identification of Bacteria
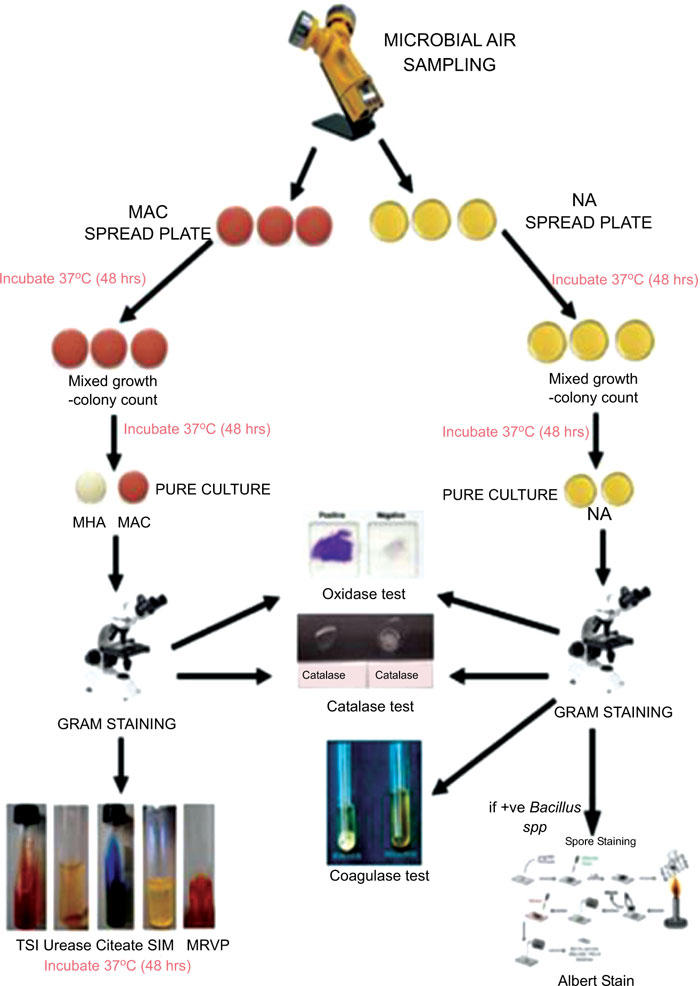
All the collected samples on Nutrient Agar (NA) plates and MacConkey Agar (MAC) plates were incubated at 37°C for 48 hours. The colonial morphology of the colonies formed was recorded, and identical colonies were sub-cultured onto NA and MAC and incubated at 37°C for 48 hours and then were proceeded for a further test such as gram staining, identification test using biochemical tests such as catalase test, oxidase test, coagulase test, Triple Sugar Iron Agar test, urease test, citrate test, Sulfide, Indole, Motility (SIM) test, Methyl Red Voges-Proskauer (MRVP) test, disc diffusion test, and Albert’s stain. Fig. (1) shows the flowchart of sampling procedures.
3. RESULTS
3.1. The Presence of Isolated Colonies in Operating Theatre
Table 1 shows that 172 (95.6%) isolates out of 180 sampling occasions in the OT were present with gram-positive bacteria, while gram-negative bacteria were isolated from 113 samples (62.8%). None of the biochemical tests were positive for Pseudomonas aeruginosa and Acinetobacter baumannii. While oxidase test for Pseudomonas aeruginosa was only positive in five (2.8%) isolates and three (1.7%) isolates were positive for Acinetobacter baumannii. Both the catalase test and the oxidase test showed a high number of positive for Bacillus spp, n=168 (93.3%), each. A further test for Bacillus spp., which was Albert’s stain, was performed for confirmation of the presence of the Bacillus spp. also showed 168 isolates (93.3%) were positive. Catalase test and oxidase test for Micrococcus spp. showed 85% of the isolates were positive (n=153). Meanwhile, for Staphylococcus spp., the catalase test and oxidase test showed a positive test for all 180 samples. However, the coagulase test showed that all 180 was negative for the presence of Staphylococcus aureus.
| Variable | Frequency (%) |
|---|---|
| Gram staining | - |
| Gram-positive bacteria | - |
| Yes | 172 (95.6%) |
| No | 8 (4.4%) |
| Gram-negative bacteria | - |
| Yes | 113 (62.8%) |
| No | 67 (37.2%) |
| Identification test for Pseudomonas aeruginosa | - |
| Biochemical test | - |
| Positive | 0 (0%) |
| Negative | 180 (100%) |
| Oxidase test | - |
| Positive | 5 (2.8%) |
| Negative | 175 (97.2%) |
| Identification test for Bacillus spp. | - |
| Catalase test | - |
| Positive | 168 (93.3%) |
| Negative | 12 (6.67%) |
| Oxidase test | - |
| Positive | 168 (93.3%) |
| Negative | 12 (6.67%) |
| Albert’s Stain | - |
| Positive | 168 (93.3%) |
| Negative | 12 (6.67%) |
| Identification test for Acinetobacter baumannii | - |
| Biochemical test | - |
| Positive | 0 (0%) |
| Negative | 180 (100%) |
| Oxidase test | - |
| Positive | 3 (1.7%) |
| Negative | 177 (98.3%) |
| Identification test for Micrococcus spp. | - |
| Catalase test | - |
| Positive | 153 (85.0%) |
| Negative | 27 (15.0%) |
| Oxidase test | - |
| Positive | 153 (85.0%) |
| Negative | 27 (15.0%) |
| Identification test for Staphylococcus spp. | - |
| Catalase test | - |
| Positive | 180 (0%) |
| Negative | 0 (0%) |
| Oxidase test | - |
| Positive | 180 (0%) |
| Negative | 0 (0%) |
| Coagulase test | - |
| Positive | 0 (0%) |
| Negative | 180 (100) |
|
Operation Room |
n | Frequency (%) | |||||
|---|---|---|---|---|---|---|---|
| Morning Session | Evening Session | ||||||
| Bacillus spp. | Micrococcus spp. | Staphylococcus spp. | Bacillus spp. | Micrococcus spp. | Staphylococcus spp. | ||
| OR ‘A’ | 9 | 7 (38.9) | 8 (44.4) | 9 (50.0) | 9 (50.0) | 7 (38.9) | 8 (47.1) |
| OR ‘B’ | 9 | 9 (50.0) | 7 (38.9) | 9 (50.0) | 7 (38.9) | 8 (44.4) | 8 (47.1) |
| OR ‘C’ | 9 | 8 (44.4) | 9 (64.3) | 9 (50.0) | 8 (44.4) | 5 (27.8) | 9 (50.0) |
| OR ‘D’ | 9 | 9 (50.0) | 7 (38.9) | 9 (50.0) | 9 (50.0) | 8 (44.4) | 8 (47.1) |
| OR ‘E’ | 9 | 8 (44.4) | 8 (44.4) | 8 (44.4) | 9 (50.0) | 6 (33.3) | 9 (50.0) |
| OR ‘F’ | 9 | 9 (50.0) | 8 (44.4) | 8 (44.4) | 8 (44.4) | 9 (50.0) | 9 (50.0) |
| OR ‘G’ | 9 | 9 (50.0) | 8 (44.4) | 9 (50.0) | 8 (44.4) | 8 (44.4) | 9 (50.0) |
| OR ‘H’ | 9 | 8 (44.4) | 9 (50.0) | 9 (50.0) | 8 (44.4) | 9 (50.0) | 9 (50.0) |
| Corridor 1 | 9 | 9 (50.0) | 9 (50.0) | 9 (50.0) | 9 (50.0) | 7 (38.9) | 9 (50.0) |
| Corridor 2 | 9 | 9 (50.0) | 7 (38.9) | 9 (50.0) | 8 (44.4) | 6 (33.3) | 8 (47.1) |
3.2. Percentage of Isolated Bacteria that Present in the OR for Morning and Evening Session
Table 2 shows the percentage of the isolated bacteria that were present in the ORs and two corridors for morning and evening sessions, which were Bacillus spp., Micrococcus spp. and Staphylococcus spp. while Pseudomonas aeruginosa and Acinetobacter baumannii were found to be absent (hence not included in the Table). The presence of Bacillus spp. was recorded 100% in OR ‘D’ and Corridor 1 as it was detectable in all nine samples for both morning and evening sessions. Similarly, Micrococcus spp. was present in all nine samples in the morning and evening sessions collected from OR ‘H’. While Staphylococcus spp. was found to be present in all nine samples in OR ‘C’, OR ‘G’, OR ‘H’, and Corridor 1 for morning and evening sessions.
4. DISCUSSION
Bacteria isolated that can be found in this study were Bacillus spp., Micrococcus spp. and Staphylococcus spp. These three types of bacteria were the most frequently isolated bacteria in all ORs and corridors. Sapkota et al. [9] observed that Staphylococcus spp., Micrococcus spp., Bacillus spp., Pseudomonas aeruginosa and Acinetobacter baumannii were the most common organisms detected in healthcare facilities.
Bacillus spp., Micrococcus spp. and Staphylococcus spp. can be transmitted from person-to-person and from air-to-people. Thus, Micrococcus spp. and Bacillus spp. were common bacteria in the environment, especially in OT [10]. Moreover, Bacillus spp. also contain spores that enable them to survive for a long period of time in the environment. Moreover, Staphylococcus spp. was usually associated with various diseases such as urinary tract infection and skin infection [11].
CONCLUSION
There were three types of bacteria isolated in this study, which were Bacillus spp., Micrococcus spp. and Staphylococcus spp. Although the bacteria identified and presence was low in percentage, but there was a chance that it can cause surgical site infection to the patients. Therefore, microbial level monitoring was very important and needs to be conducted as prevention measures.
Recommendation to maintain or reduce bioaerosol level in the OT can be made by strengthening surveillance of the hygiene in the OT. This can be achieved by making sure the ventilation systems provided in the OT was well checked so that it can generate adequate ventilation to reduce microbial in the air of OT. In addition to that, it was improving a hygiene procedure such as sweeping, regular cleaning, and disinfection of the hospital appliances in all health care settings before and after surgery was done. Overcrowding in the OT should also be avoided in the OTs because crowded places can increase the chances of infectious disease. Lastly, more effort is needed to improve the OT hygienic condition and recommendation by giving awareness and educational knowledge on how to reduce the hazard of airborne transmission, which was potentially pathogenic microorganism that can cause nosocomial infection toward the patients.
It is also suggested that in the future, the study can be done by using a swab sampling method for hospital appliances that are placed in the OT. It is also recommended that additional measurements such as the ventilation rate and fungal load were conducted, then further checking on the strains to be performed using the Polymerase Chain Reaction.
LIST OF ABBREVIATIONS
| CDC | = Centre for Disease Control and Prevention |
| HAI | = Hospital-Acquired Infections |
| HUSM | = Hospital USM |
| MAC | = MacConkey Agar |
| MRSA | = Methicillin-Resistant Staphylococcus aureus |
| MRVP | = Methyl Red Voges-Proskauer |
| NA | = Nutrient Agar |
| OR | = Operation Room |
| OT | = Operating Theatre |
| SIM | = Sulfide, Indole, Motility |
| SSI | = Surgical Site Infection |
| WHO | = World Health Organisation |
ETHICS APPROVAL AND CONSENT TO PARTICIPATE
Not applicable.
HUMAN AND ANIMAL RIGHTS
No animals/humans were used for studies that are the basis of this research.
CONSENT FOR PUBLICATION
Not applicable.
AVAILABILITY OF DATA AND MATERIALS
The data that support the findings of this study are available from the corresponding author [S.M.A] on request.
FUNDING
None.
CONFLICT OF INTEREST
The authors declare no conflict of interest, financial or otherwise.
ACKNOWLEDGEMENTS
The authors are grateful to Hospital USM, Associate Prof. Dr. Siti Suraiya Md Noor and all staff at the CDC Unit and OT Department, postgraduate student of Biomedicine (Nanda), laboratory staff of Biomedicine Program (Siti Kurunisa Mohd Hanafiah and Muhammad Syafiq Sidek Ahmad), laboratory staff of Environmental and Occupational Health Program (Mrs. Juskasmini Jusoh and Mr. Md Khairul Azuan Bin Che Azid), as well as Siti Nur Syamimie Zamanie and Nur Atiqah Hanim Lukman, who have been extremely helpful.


